Extract-N-Amp™ Tissue Feature
Scott Weber, Derek Douglas, St. Louis, MO, USA
Introduction
Standard methods for extracting DNA from tissues can be extremely laborious and time consuming. Certain applications, such as genotyping of transgenic mice using a section of tail, employ a lengthy DNA extraction process. For example, traditional mouse tail DNA extractions require a 5 hour to overnight digestion with proteinase K and subsequent phenol:chloroform extraction and alcohol precipitation. Recently other options such as silica bind and elute kits have become available, but these also require a 5 hour to overnight digestion of the tail before the DNA is extracted. These time consuming and labor intensive methods are necessary for some applications, but for more robust techniques, such as PCR, the DNA does not need to be isolated to the degree that these methods provide.
PCR can amplify a target using very little template and from relatively impure starting material. Therefore, the Extract-N-Amp™ Tissue PCR Kit has been created to release PCR-ready DNA from mouse tails in a 15 minute single tube procedure. The included 2x PCR mix is optimized to work with the crude extracts and the solutions in the kit.
We describe a new method from which sufficient DNA for PCR amplification is released from a mouse tail after a 10 min extraction. The neutralized tissue extract is stable for at least 6 months at 4 °C, and can be added directly to the optimized PCR mix which is supplied in the kit. Furthermore, the Extract-N-Amp™ Tissue Kit works with a variety of source materials including fresh and frozen mouse tails, mouse tissue, buccal swabs, hair, saliva and saliva archived on a collection card.
Materials and Methods
All materials were obtained from St. Louis, MO unless otherwise noted. All PCR primers were obtained from Sigma-Genosys (The Woodlands, TX)
DNA extraction
Extraction Solution and Tissue Preparation Solution were mixed together and tissue samples were added to the mixture (Figure 1). The mixture was incubated at room temperature for 10 minutes to extract the DNA. An incubation at 95 °C for 3 minutes stopped the extraction and Neutralization Solution B was then added to neutralize the extract. Samples were mixed thoroughly and used immediately in PCR. The volumes of the solutions used and incubation times for each starting material are listed in Table 1.
PCR amplification
For standard PCR, 10 µL of REDExtract-N-Amp™ PCR ReadyMix was combined with forward and reverse primers at 0.4 mM and 4 µL of neutralized tissue extract in a final volume of 20 µL Reactions were assembled at room temperature. Cycling was performed in a GeneAmp® PCR System 9700 thermal cycler (Perkin Elmer/Applied Biosystems, Foster City, CA). The PCR primers used for each sample type and the cycling parameters used are listed in Table 1. The sequences for the primers are presented in Table 2. Aliquots of 3-6 µL of each PCR product was loaded and fractionated on an ethidium bromide-stained 1% agarose gel in Figures 2 and 3.
For quantitative PCR, 10 µL of Extract-N-Amp™ PCR ReadyMix was combined with forward and reverse primers at 0.4 mM, SYBR® Green I dye (S9430), 0.6 µL of Reference Dye for Quantitative PCR (R4526), and 4 ml of tissue extract in a final volume of 20 ml. Quantitative PCR was performed in an ABI PRISM® 7700 Sequence Detection System (Perkin Elmer/Applied Biosystems, Foster City, CA). Primers and cycling parameters used are listed in Table 1.
Mouse DNA standards for quantitative PCR were prepared from 1.2 cm of mouse tail using the GenElute™ Mammalian Genomic DNA Kit (G1N10, G1N70 or G1N350). The purified DNA was diluted to 57.5 ng/ml and seven 2-fold serial dilutions were prepared from the initial dilution. Single-use aliquots of these DNA standard dilutions were stored at -20 °C. A set of standards was assayed along with the mouse tail extracts.
Sequencing
PCR products prepared from mouse-tail extracts with the IL-1β primer pair were sequenced after purification with the GenElute™ PCR Clean-up Kit (NA1020). The sequence was obtained using BigDye™ Terminator Chemistry on an ABI PRISM® 377 DNA Sequencer (Applied Biosystems, Foster City, CA).
Results and Discussions
PCR amplification from tissue extracts
The Extract-N-Amp™ Tissue PCR Kit was used to extract and amplify target DNA from mouse tails, mouse brain tissue, buccal swabs, human hair, saliva and saliva archived on a collection card. As shown in Figure 2, PCR products ranging from 1-2 kb were detected on ethidium bromide stained 1% agarose gel from each extract. The data in Figure 2 demonstrates that PCR-ready DNA can be rapidly extracted from a variety of tissue sources. Additionally, experiments are in progress using the Extract-N-Amp™ Tissue PCR Kit to extract DNA from other starting materials such as Drosophila and fungi.
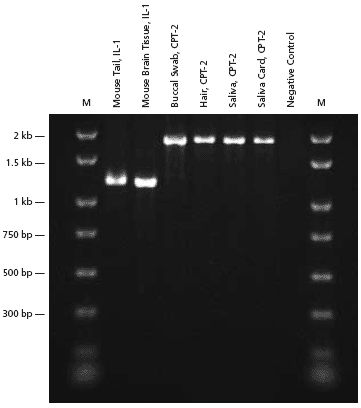
Figure 1.PCR analysis of genomic DNA extracted from various types of sources.
Extract-N-Amp™ Tissue PCR Kit was used to extract and amplify genomic DNA from various sources. Genomic DNA was extracted using the protocol as described in Materials and Methods. In all cases, the entire extraction procedure was completed in less then 15 minutes. All samples were then amplified using the specially formulated Extract-N-Amp PCR ReadyMix™ included in the kit. PCR products generated are 1181 bp for the Interleukin-1β (IL-1β) gene in mouse and 1820 bp for the Carnitine palmitoyltransferase II (CPT-2) in human.
It is known that both the reaction conditions and the template quality determine the size of PCR product that can be amplified.1 Using the Extract-N-Amp™ Tissue PCR Kit a 5 kb fragment of β-Globin (Figure 2) was successfully amplified using extracts from buccal swab, hair, saliva and saliva archived on a collection card. The results shown in Figures 1 and 2 demonstrate that reaction conditions and template quality obtained with the Extract-N-Amp™ Tissue PCR Kit are suitable for amplification of both small (< 2 kb) and medium (5 kb) sized PCR products.
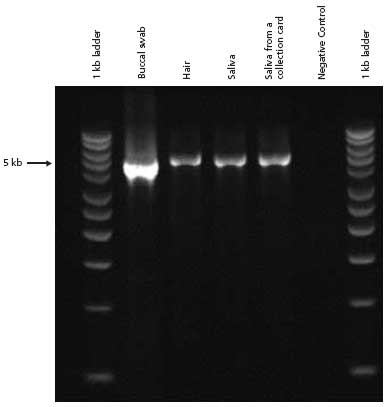
Figure 2.PCR analysis of genomic DNA extracted from various types of human tissue and amplified with 5 kb β-Globin primers. Extract-N-Amp™ Tissue PCR Kit was used to extract and amplify genomic DNA from various human tissue sources.
Genomic DNA was extracted from buccal swabs, hair, saliva, and saliva archived on a collection card using the protocol as described in Materials and Methods. All samples were then amplified using the specially formulated Extract-N-Amp PCR ReadyMix™ included in the kit. The PCR products generated are 5 kb for a segment of the Human β-Globin gene.
It should be noted that the Extract-N-Amp™ Extraction Solution contains components that are not compatible with standard PCR mixes. The 2x Extract-N-Amp™ PCR Mixes are specifically formulated to compensate for these components and other impurities released during the extraction process. Extracts added to a PCR mixture other than the Extract-N-Amp™ or REDExtract-N-Amp™ PCR ReadyMix will likely yield little or no PCR product.
Stability of mouse tail extracts
To test the stability of the DNA extracts prepared with the Extract-N-Amp™ Tissue PCR Kit, stability tests were set up with mouse-tail extracts utilizing quantitative real time PCR. Four mouse tails were extracted according to the procedure outlined in the Materials and Methods section and 4-µL aliquots were analyzed immediately by quantitative PCR with SYBR® Green dye detection on an ABI PRISM® 7700. The Extract-N-Amp™ Tissue PCR Kit extracts are amplified without significant quenching of the SYBR Green signal. As seen in Figure 3, the amplification plots for the extracts fall in the middle of the amplification plots for the genomic mouse DNA standards (Figure 4).
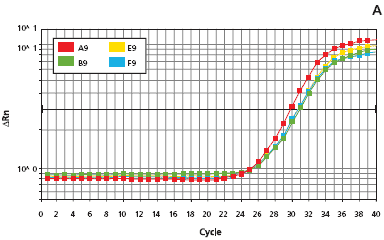
Figure 3.
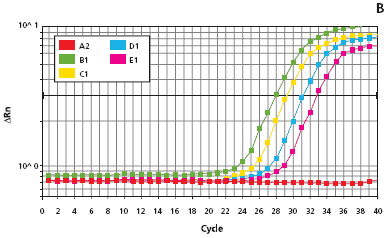
Figure 4.Quantitative PCR results showing mouse tail extract stability. Four mouse tails were extracted using the Extract-N-Amp Tissue PCR Kit following the procedure described in Materials and Methods. Plot A shows the amplification of four mouse tail extracts immediately following extraction. Plot B shows the amplification of the standards used to quantify the DNA in the mouse tail extracts. Reaction analyses were performed on an ABI Prism™ 7700® Sequence Detection System.
Half of the remaining extracts were stored at 4 °C (recommended storage conditions) and the other half at 37 °C (accelerated storage). Quantitative PCR was performed after 5 and 10 weeks for all extracts and on separate aliquots of the same standards used at time zero. As shown in Figure 5, mouse-tail extracts are stable at 37 °C for 10 weeks. Results for extracts stored at 4 °C are very similar to those shown in Figure 5 (data not shown). It should be noted that the mouse tail was removed from the neutralized extract before storage.
Results from stability experiments performed on the mouse tail extracts using the Extract-N-Amp™ Tissue PCR Kit indicate that DNA will be detectable beyond the recommended storage time of 6 months at 4 °C. Additionally, experimental data suggests that the extracted DNA should be detectable up to 12 months after the original extraction.
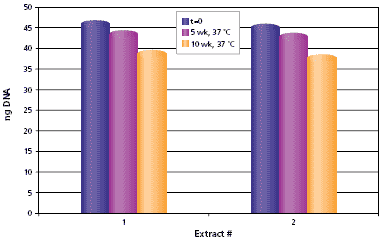
Figure 5.Stability of DNA in mouse tail. Two mouse tails were extracted using the procedure described in Materials and Methods. Mouse tails were removed from samples prior to storage. The extracts were stored at 37 °C (accelerated) and quantitative PCR was repeated after 5 and 10 weeks with the extracts.
Sequencing of PCR products from mouse tail extracts
PCR products produced with the Extract-N-Amp™ Tissue PCR Kit can be sequenced directly, however higher quality sequence is often obtained after purifying the product through a routine clean-up procedure. As illustrated in Figure 5, a clean trace was obtained from mouse-tail extracts by sequencing with the original primers (IL-1b) used for PCR. Prior to sequencing, the PCR product was cleaned up using the GenElute™ PCR Clean-Up Kit (NA1020). The nucleotide sequence was checked against the published sequence (NCBI Accession Number: X04964) and found to be accurate.
Conclusions
The Extract-N-Amp™ Tissue PCR Kit extracts DNA suitable for PCR in 15 minutes, using a straightforward protocol. While most commonly used with mouse tails in genotyping experiments, the kit has been shown to work with a variety of starting materials, including mouse tissues, buccal swabs, hair, saliva, saliva archived on a collection card, and C. elegans. DNA prepared with the Extract-N-Amp™ Tissue PCR Kit is stable for at least 6 months at 4 °C, allowing for multiple reassays. Short to medium-sized PCR targets (up to 5 kb) can be amplified from the extracts on a consistent basis. Furthermore, the PCR products are suitable for sequencing.
Acknowledgments
The authors thank Jessica Copeland, Jaime Miller, and Bruce Blackerby of R&D for providing the sequence data, Kevin Ray and Carol Kreader for their critical reading and contributions to the preparation of this article. Also thanks to Dave Hanson and Sudhir Nayak, from the Tim Schedl lab at Washington University, St. Louis, MO.
Refer to Table 1 and Table 2 below for PCR primers used for each sample type, cycling parameters, and sequences for the primers.
Ordering Information
- NCBI Accession Number: X04964
- NCBI Accession Number: U01317
- Wormbase gene designation F10B5.7
References
To continue reading please sign in or create an account.
Don't Have An Account?