Validating CRISPR/Cas9-mediated Gene Editing
T7E1 Assay Versus Sequencing Methods and When to Use Them
Conducting a CRISPR experiment involves many steps, with each of these having options to weigh. After choosing a target gene, designing the appropriate guide RNAs (gRNAs), and ordering the guides and necessary reagents, the next stage is validating that the guides have successfully targeted the gene of interest and enacted the desired change in expression (Figure 1).

Figure 1A.Design gRNA, choose Cas nuclease and select transfection method.
Figure 1B.Introduce CRISPR components using selected transfection method to edit targeted genes in desired cell line.

Figure 1C.Analyze gene editing results to confirm accuracy and efficiency via T7E1, Sanger sequencing, NGS, Western/Elisa, FACS
Here, we give an overview of available validation methods including both their mechanisms and relative merits to help inform your workflow choices.
The Importance of Controls in CRISPR Validation
Because validation is a such an important part of a gene editing experiment, having the proper controls is vital, providing the basis for sound analysis. A typical negative control consists of a gRNA that does not target any known sequence in your experimental system and is introduced into cells with all of the same reagents as the gRNA targeting your gene of interest. This allows any observed phenotype to be attributed to the specific effects of gene loss of function rather than any off-target effects of the reagents used to generate the knockout.
Consisting of at least one, pre-validated, high-efficiency gRNA under identical experimental conditions to the gRNAs being validated, positive controls are also an integral part of any gene editing experiment. Often, housekeeping genes are used for this purpose. Positive controls are particularly important when a lack of cutting is observed to ensure it is not a result of an error in the experimental procedure.
Assessing CRISPR Targeting
There are two categories of commonly-used methods to detect whether a given gene has been successfully edited in a CRISPR experiment. The first type exploits the ability of enzymes to target and cleave mismatched DNA to test for the presence of mutant alleles. The second type is sequencing-based, allowing direct assessment of the DNA itself. Both types of validation can be useful at different parts of a CRISPR workflow.
Enzyme Mismatch Cleavage Assay for First-pass CRISPR Validation: T7E1 Assay
| Initial Denaturation | 98 °C | 30 Seconds | x1 |
| Denaturation | 94 °C | 15 Seconds | x35 |
| Annealing | 60 °C | 30 Seconds | x35 |
| Extension | 68 °C | 45 Seconds | x35 |
| Final Extension | 68 °C | 5 Minutes | x1 |
Enzyme mismatch cleavage (EMC) techniques are generally inexpensive and easy to conduct with standard laboratory equipment, so are often used to screen cells for successful gene editing. T7 endonuclease I (T7E1)1 is a bacteriophage-derived mismatch-specific enzyme commonly used for CRISPR validation and can be found in the Sigma-Aldrich® T7E1 kit.
The first step of T7E1-mediated validation is to harvest genomic DNA and amplify the region surrounding the gRNA target site by PCR (Table 1). If a mutation was successfully introduced into one allele by non-homologous end joining (NHEJ) after CRISPR/Cas9-mediated cleavage, this results in amplification of both wild-type and mutant sequences. A high-fidelity DNA polymerase like the AccuTaq™ enzyme, is critical at this stage in order to prevent false positives as a result of mutations introduced in the amplification step rather than by CRISPR-mediated gene editing.
After amplification, the resultant DNA is heated and cooled to denature and reanneal it, generating some products that are heteroduplexes of wild-type and mutant amplicons, some entirely wild-type, and some entirely mutant. All of these are then incubated with T7E1 enzyme, and the reaction products run on an agarose gel for analysis (Figure 2). The mismatched sequences of heteroduplexes change DNA structure, making them a target for cleavage by T7E1 and generating a specific and predictable size of cleavage products based on PCR primer and gRNA binding sites. The intensity ratio of cut vs. uncut bands on an agarose gel can be used as a readout of gene editing efficiency.
Validation Process Overview
Gene editing experiment validation using T7E1

Step 1
Isolation of Genomic DNA - Modified Iocus

Step 2
PCR Amplification

Step 3
Denaturation & Annealing - Possible re-annealing products

Step 4
Digestion of Mismatched Duplexes - T7 Endonuclease I

Step 5
Agarose gel electrophoresis
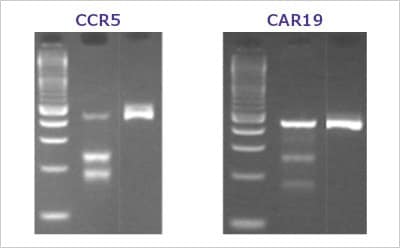
Figure 2. A typical T7E1 assay resultHEK293 cells were transfected with various Cas9 and sgRNA RNP complexes targeting the CCR5 and CAR19 genes. After three days, transfected cells were harvested and the genomic DNA was collected. The target genes were PCR amplified using AccuTaq™ LA DNA polymerase (D8045) and the editing analyzed using the T7 Endonuclease I Gene Editing Detection Kit. Lanes are ordered by ladder, sgRNA edited cells, negative control.
The simplicity of the EMC workflow has made it a popular method for CRISPR validation. It is easy to analyze, providing same-day results with no requirement for sequencing or complex software. However, this method cannot determine the specific mutations, and can generate false positives as a result of naturally-occurring polymorphism.
Sequencing-based Methods for CRISPR Validation
While EMC methods can perform an effective first-pass analysis of gene editing in a CRISPR workflow, those results often must be confirmed with sequencing to ensure that the desired mutation was introduced. This is a necessary step because a gRNA can show strong cutting in an EMC assay, but may not introduce a mutation useful for protein knockout because, for example, it is in-frame and produces a protein that may retain function. While it may seem that using sequencing as an initial screening tool would be beneficial because it is more specific than EMC analysis, the higher cost and longer turnaround prevents sequencing from being used in this way.
Sanger Sequencing and Tracking of Indels by Decomposition (TIDE)
Sanger sequencing is a traditional and popular technique for gene editing validation.3 In this method, DNA is amplified using a specialized type of PCR called chain-termination PCR where a low concentration of fluorescently-labeled, modified nucleotides that terminate DNA extension—dideoxyribonucleotides (ddNTPs)—are mixed with dNTPs to generate DNA chains of every possible length. Using a different fluorophore for each of the bases allows determination of the sequence after separating the mixture of DNA chains by size using capillary gel electrophoresis.
Sanger sequencing is very reliable, but has drawbacks. First, it is time-consuming as a result of the many experimental steps. It also is not amenable to high-throughput analysis, as it only sequences a single DNA fragment at a time. In general, this is not an issue for validation of successful editing of a single gene, but does not allow for determination of off-target effects in the same reaction.
TIDE is an analysis method based on Sanger sequencing where a mixed population of edited and unedited cells is compared by a software algorithm to reveal the presence of indels and their frequency.4 It is a quick, sensitive, and cost-effective way to find mutations that does not require a clonal population of cells to conduct, but the nature of the algorithm prevents sequence identification without more detailed methods.
Next Generation Sequencing (NGS)
NGS is a highly sensitive method capable of identifying low-frequency mutations in a cell population.3 Like Sanger sequencing, it analyzes DNA fragments, but differs in that it does so with massively parallel sequencing technology for much higher throughput. Because it can analyze so many fragments in parallel, it can be used to assess for off-target effects in CRISPR experiments. While it has important advantages over other validation methods, it is also error-prone and is the most costly technique discussed here.
Confirming Loss of Expression
In addition to validating successful gene editing by confirming the DNA sequence, it is also critical to confirm that the desired modification occurred in the protein product. When evaluating a knockout, introduction of insertions and deletions does not necessarily lead to complete disruption of expression: some genes may still produce truncated or partial proteins. Confirmation can be conducted at the mRNA level using techniques like RT-PCR for evaluating accurate integration of mutations or SNPs, while confirmation of gene knockout and addition of a fusion element such as GFP should be confirmed by gel electrophoresis and Western blot using one or more validated antibodies.
Assessing CRISPR/Cas9 Gene Editing Isn’t One-Size-Fits-All
Conducting CRISPR/Cas9-mediated gene editing is complex and should be tailored to meet the goals of your study. As a result, strong controls and appropriate validation of both gene editing and loss of expression are critical to success. Number of samples, cost, turnaround time, and type of results are all factors that will influence your choice of validation method (Table 2).
| Method | Readout | Recommended Uses | Specialized Equipment | Bioinformatic Analysis Required | *Turnaround Time | Cost |
|---|---|---|---|---|---|---|
| T7E1 | Rough indel size and efficiency estimate | Optimization of experimental design; evaluation of several gRNA sequences | No | No | Same day | $ |
| Sanger | Detailed sequence information at target site | Confirmation of T7E1 results with detailed sequence information | Yes | Yes | 2-14 days | $$ |
| NGS | Detailed sequence information including off-target sites and frequency | Large number of samples; Evaluation of off-target editing; Ideal for preparing regulatory reports | Yes | Yes | 3-5 days | $$$$ |
Determining the appropriate balance of all of these factors can be challenging. If you would like to get more information or discuss the options that fit your needs, speak to one of our experts. Our team will support you through every step of your gene editing experiments.
Related Products
Response not successful: Received status code 500
References
To continue reading please sign in or create an account.
Don't Have An Account?